Research OverviewCellular viability and function require exquisite spatial organization and temporal coordination over a broad range of length- and time- scales. At the nanoscale, proteins must be allowed to explore appropriate ensembles of conformations in order to guarantee the right interactions with their environment and with each other, and yet, this level of organization would be meaningless were it not also the case that the overall spatial distribution of proteins and other cellular components is tightly regulated at the microscale. Moreover, events at every scale are strongly coupled: a single mutation in a protein sequence can lead to misfolding, aggregation, and inclusion formation that is ultimately visible under a microscope. The coalescence of such an inclusion can, in turn, so dramatically alter the load experienced by cellular protein folding quality control machinery that the conformational states of many proteins may be altered. Until recently, limitations in theoretical, computational, and experimental methods prevented researchers from investigating systems of interest on multiple scales simultaneously: scientists either attempted to crystallize a fragment of a single protein, or observed cellular behavior in ignorance of many events occurring at the molecular level. Simultaneous advances in a variety of technological areas now allow significant new discoveries in the field of cell biology to be achieved. Foremost among these, super-resolution fluorescent microscopy has been adapted to the examination of protein interactions in live cells with molecular-scale precision. For the first time, it is possible to correlate genuinely biological phenomena at the level of the whole cell with specific molecular events that cause them, shifting the fundamental paradigm of molecular biology out of the test tube and into the cell where life takes place.
Our goal is to provide the first complete spatial map of protein folding as it takes place in vivo, focusing on cell culture and yeast models of neurodegenerative disease as systems for studying protein folding quality control. The many different pathways -- such as synthesis, ubiquitination, or interaction with chaperones -- that regulate and modulate folding have been extensively characterized in vitro, and yet in order to comprehend the coordinated action of these different components of the cellular machinery, it is essential to observe how they combine to form a dynamic and highly organized quality control super-structure that extends throughout the cell. The dynamic compartmentalization of a whole host of biochemical processes (ranging from protein translation and degradation to RNA quality control and viral assembly) remained invisible to conventional biochemical methods, which do not inspect the cell optically before lysis and homogenization. Now, for the first time, imaging techniques such as super-resolution imaging and laser manipulation have emerged as indispensable tools for resolving various distinct populations of macromolecules and correlating each molecule's trajectory through space with biochemical interactions and changes in conformational state. The application of new high-resolution, dynamic imaging technologies enable us to relate biochemical and genetic information to a spatio-temporal map of events in the process of cellular protein folding quality control. |
Projects
The protein folding problem
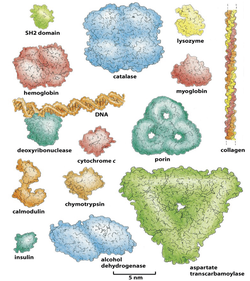
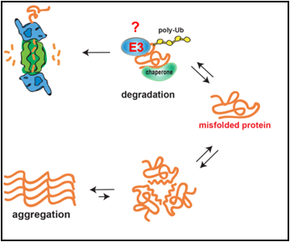
The common denominator of all proteins in the cell is their need to maintain a set of low-energy conformations of their folding structure, known as the native state. Folding to the native state is essential to protein function, and proteins that fail to do so pose a threat to the cellular environment by exhibiting a tendency to misfold, aggregate, and otherwise disrupt the cellular environment. Roughly speaking, folding to the native state involves the burial of hydrophobic amino acids inside the core of the protein, and the exposure of hydrophilic amino acids on the outside of the protein. Our understanding of the relationship between amino acid sequence and protein structure are complicated, however, by several factors:
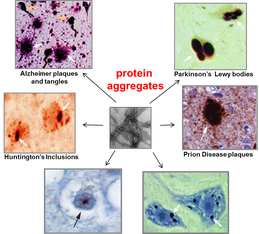
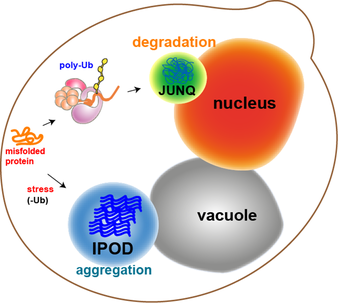
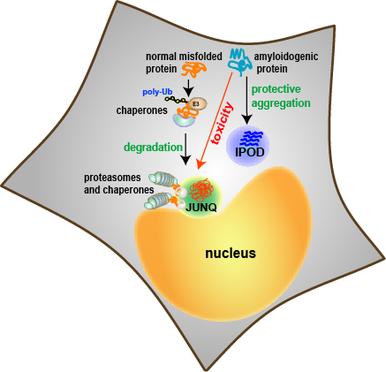
1. proteins in the cell are not in an ideal aqueous environment - crowding effects and local nano-environments will tend to dominate thermodynamic considerations involved in folding. 2. protein function and protein-protein interaction is driven by hydrophobic residues that are not buried in the core of the protein. 3. protein structure is dynamic - even in the native state a protein may sample multiple conformations. What interests us is the question: what is the difference between a native protein and a misfolded protein from the standpoint of the cell and its protein folding quality control machinery? We approach this problem in collaboration with the lab of Jeremy England, who has recently developed a ground-breaking theoretical bio-physical model for protein misfolding. With out cell biological tools and in vivo models of protein misfolding, we can now apply Jeremy's theoretical approaches to protein misfolding problems in the cytosol of a living cell. Our joint aim is to understand the following questions: How does the cell recognize a protein as being misfolded (as opposed to a natively unstructured protein, a protein that has a dynamic structure with multiple allosteric conformations, or a completely unfolded protein)? How does the cell distinguish between misfolded proteins that should be degraded or refolded and amyloidogenic proteins that have no hope of folding and should be taken to an aggregate inclusion? |
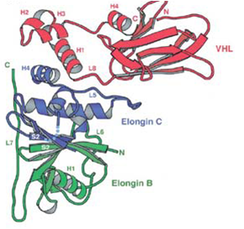
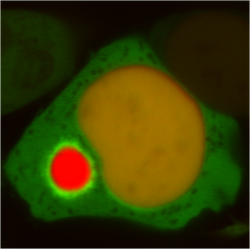
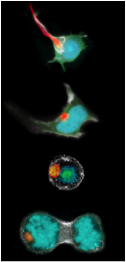
The von-Hippel Lindau protein (VHL) is a tumor suppressor that has been extensively studied as a model for protein misfolding. The VHL protein (red) is constitutively misfolded unless it can bind to its binding partners Elongin B/C (blue and green). Several tumoragenic mutations disrupt the binding and lead to VHL misfolding and degradation by the protein folding quality control machinery of the cell.
In collaboration with the lab of Jeremy England, we are studying the relationship between VHL sequence and its tendency to misfold. Our first results from this project were published in Brock et al., Structure 2015. |